A kind of magic
Bacterium or blue whale, every living thing leaves a trace. Now we have a tool that can find that trace, in soil, in water—even in the air—and it’s changing the way we do science.
Shaun Lee hunts a ghost, a silvery apparition that dwells now only in museums and regret. And maybe—just maybe—a last few backwaters. The grayling is the bad-joke poster child of Aotearoa’s freshwater management: famous as our only protected freshwater fish, and farcical for the fact that it’s extinct.
European settlers arrived to find rivers boiling with grayling. In 1871, eminent naturalist Sir James Hector wrote that they were found in “most of the streams of the colony” and described “immense shoals” swimming up Wellington’s Hutt River that summer. But something went wrong for the grayling, and experts think it was almost certainly the release of brown trout in 1867. Such was the carnage that within a decade, Nelson amateur naturalist Joshua Rutland reported that grayling had already disappeared from the Maitai River.
The last time anybody indisputably saw one was in 1925, in the Waipu River in Northland, although newspapers ran anecdotal reports of sightings up to the 1950s. Freshwater ecologists who’ve modelled its oblivion say a few may have hung on in the backcountry until the 1970s. But Lee reckons it’s worth one last look.
His chances, he allows, are as remote as the distant reach of Lake Alice, at the head of Fiordland’s George Sound. But this is where the experts pointed him, on account of the surging waterfall that is the lake’s only connection with other waterway networks. Late in March 2022, as the helicopter came into land, Lee eyed that waterfall with relief. “It looked steep enough to hold back brown trout.”
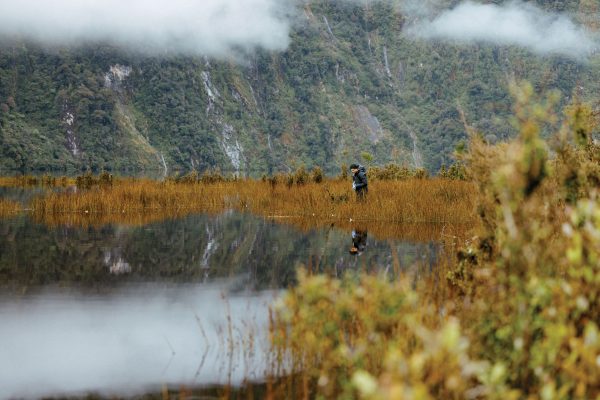

There was but one small landing option, and it was boggy. Lee would have to work fast, while the pilot waited. The trip had cost him as much as a trip to London.
“I felt guilty about taking a helicopter,” he says, “but my window for sampling that season was closing.” They’d left Te Anau early that morning, turned with the rising sun over the Murchison Mountains. For Lee, there was something auspicious in that: “I’ve tracked takahē in there, so I know that things sometimes come back from the dead,” he says.
“I’m pretty sure we’ve lost the grayling, but there’s a small part of me that says ‘maybe’, and that’s enough to motivate me.” Lee, an Auckland photographer, was born in 1975, about the time it’s thought the last grayling were finally dying out, “so I feel like there’s still a sort of connection there—it doesn’t feel too distant, too impossible”.
After all, it’s not like he has to actually catch a grayling, or even see one. All he has to do is fill a few small syringes with water.
[Chapter Break]
As I sit at my desk, I watch a flurry of airborne dust waft across a window, set asparkle by a low winter sunbeam. Many of those glittering particles were once me.
You might shed as much as a gram of your skin—200 million cells—in the next hour. By your last day, 35 kilograms of old skin will litter the path of your life, and under the right conditions, some of it will still hold traces of your DNA. Once it’s left your body, geneticists call it “eDNA” (the e stands for “environmental”), and if it fell indoors within the last nine or 10 months, they could probably still obtain your entire genetic profile from it.
That’s thanks to sophisticated eDNA analysis, developed only in the past two decades, that can pinpoint the genetic signature—RNA or DNA—of practically any organism. You just need an existing sample to compare yours to. The first step is to extract DNA/RNA from the cells you’ve collected, be they skin, fur, feathers—or often, as in the case of COVID wastewater testing, poo. Jo Chapman, senior scientist at the Institute of Environmental Science and Research, explains the process. Clumps of cells are sometimes shaken apart, or ground like coffee beans. The mash is washed in a detergent, then in protein enzymes, to dissolve most of the unwanted material. Finally, it’s filtered. Then the DNA/RNA is brought back from solution into discernible form by adding ice-cold alcohol.
If the genetic signature you’re looking for is there at all, it’s in infinitesimal amounts—not enough to detect. So scientists feed the sample to a machine that’s essentially a genetic photocopier.
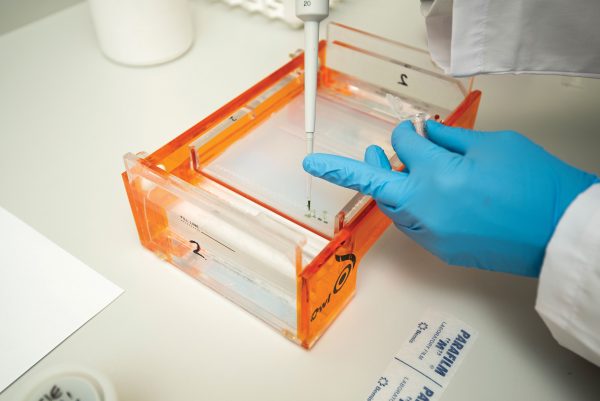


Most organisms—you included—carry a genetic code “written” in just four characters, A, C, G and T. They’re shorthand for the bases—adenine, cytosine, guanine and thymine—that comprise a molecule of DNA. The order they’re arranged in determines the identity of the organism, and the next machine, a sequencer, can read that order, much as the scanner at a supermarket checkout identifies a product by its barcode. “Now you have the unique combination of As and Cs and Gs and Ts,” explains Chapman, “which means you can work out what that organism is.”
Wastewater testing is very specific, she says. “We’re looking for COVID-19 and nothing else.” But eDNA sampling has much broader applications, and they’re helping scientists find evidence of organisms in all manner of places.
Under the right circumstances, DNA can be incredibly hardy: geneticists have recovered samples of animal and plant eDNA from soils in northern Greenland that are two million years old. There are genetic echoes of organisms everywhere—in water, sediment and soil; researchers can even pull eDNA out of thin air. In 2022, Canadian geneticist Elizabeth Clare placed 70 air pumps and filters around the Hamerton Zoo Park in England. It was, she told reporters, “a bit of a crazy idea—vacuuming DNA out of the sky”.
But why a zoo? One of the biggest challenges of working with airborne eDNA, Clare explained, is avoiding contamination from other sources, which could mess up results. “My lab doesn’t handle tiger DNA ever, so if we’re detecting a tiger, there is no other source.”
When she headed back to the lab, Clare got her tiger, plus another two dozen species known to be held at Hamerton. Her filters even detected DNA from the food the animals were given. Momentously, her devices also detected hedgehogs outside the fence, more than 200 metres away. For scientists such as Susie Wood, a molecular ecologist at Nelson’s Cawthron Institute, that keen perception is manna from heaven, because creatures like hedgehogs—secretive, nocturnal or cryptic—often evade conventional surveys. “It’s an incredible tool,” she says. “It’s totally revolutionised how we do environmental biology.”
There are two basic approaches to eDNA work. One is the targeted method used to look for COVID in wastewater. Wood explains: “We might take a sample that contains a soup of DNA from many different organisms, but we look specifically for the species in question.” The other approach is meta barcoding, or community profiling, where you catalogue all of the biodiversity in a sample. Say you want a list of all the plant species in a certain stream: community profiling is the way to go.
This approach was developed back around 2010, Wood says. “But it was very expensive back then, and we were getting huge volumes of data. We had to get our heads around how the computing power might cope. Now we can do it on a laptop in half an hour.”
Cawthron has built a dedicated multimillion-dollar eDNA lab, and it’s already turning up surprises.
Water sampling recently found a new population of the world’s rarest kākahi, or freshwater mussel, in a dune lake in Golden Bay/Mohua.
Echyridella onekaka wasn’t even described until 2006. It’s thought to live only in north-west Nelson, and ecologists know almost nothing about it. “We weren’t expecting to find it there,” says Wood, “but we had positive results from all seven samples, so we were very confident in our results. It’s super rewarding to find such a rare species in a new location.” And it wouldn’t have happened without eDNA: “There was an intense cyanobacterial bloom at the time, so divers would probably have missed it.”
Wood shows me through the new lab. Each step of the process happens in a different room, and all of them are bathed in UV light to kill off any stray material that would confound the result. “The technique is incredibly sensitive,” she says, “so we have to be very careful.” By the final room “we have a genetic list of species, but they’re still just As and Cs and Gs and Ts. So then we compare our sequences to an international reference database, and that spits out a name, be it pigeon or kākāpō or moa.” Or it might draw a blank. Of the estimated 8.7 million species on the planet, just over a million have been described, and still fewer have had their DNA committed to record. “We haven’t got a barcode for every species in the world.”
That means Wood must interpret results carefully. “Some things are very similar, genetically, and if the database doesn’t have an exact match, it’ll give you the nearest matching sequence. That can be something completely different. We once got a result back from a lake sample that said the closest match to one sequence was the eastern grey gorilla. We had a lot of fun joking with each other about whose DNA might have contaminated the sample.”
[Chapter Break]
Fortunately for Shaun Lee’s grayling hunt, the global database has plenty of extinct species on it. It took years, but Otago University geneticists managed to sequence the grayling’s genome from the 23 specimens that remain here and in the United Kingdom. Then they sent it to Shaun Wilkinson, founder of Wellington eDNA facility Wilderlab, who compared that grayling marker with results from thousands of old eDNA samples in his database. Says Lee: “Basically, he did the biggest-ever backwards search for an endangered species in New Zealand on his laptop, and that’s the magic, the power of eDNA. I had no idea it could work in reverse.”
Wilkinson drew a blank, but Lee was undeterred. He pored over maps, looking for rivers and streams brown trout couldn’t get to. “There aren’t many of those left,” he says. Grayling used to migrate each spring to the sea, but Lee thinks some landlocked lakes might still be contenders. “If they’re fed by upland streams, the fish might spend the winter months in headwaters, then come down and use the lake as a proxy ocean in summer.”

And so it was that he ended up at Lake Alice, racing to fill little syringes before his helicopter sank into the mire. Meanwhile, he says, “the sandflies were sampling me”. DNA can be patchy in slow water—a single sample might easily miss that precious trace of grayling—so Lee had to plug about the marshy lake edge, spacing out his best-practice six replicate samples. Despite the hustle, he took a moment.
It was a stunning morning—still, pristine, mist wrapped around the ridges. “The whole catchment looked untouched—a reminder of what Aotearoa was like before we changed it all forever. It was exactly what hope should look like, and I did feel hopeful.”
[Chapter Break]
For ecologists, eDNA’s extraordinary sensitivity can bring blessing and bafflement. In 2021, samples by Wilderlab from the sea off Breaker Bay returned positive for kororā/little blue penguin, stingray, orca—and eastern grey kangaroo. At times like this, researchers have to think laterally. The samples also contained a smoking gun: dog DNA. A bit of detective work established that a nearby supermarket sold dog food containing kangaroo meat. In 2022, analysis found capybara DNA in Canterbury’s Pūharakekenui/Styx River. Capybara are a giant, water-loving cavy found in South America but also, it turned out, in the Willowbank Wildlife Reserve in North Christchurch. By way of compensation, the survey also found the DNA of pekapeka-tou-roa/long-tailed bats. While they’re still holding on around Geraldine, they’d not been recorded in Christchurch since 1885.
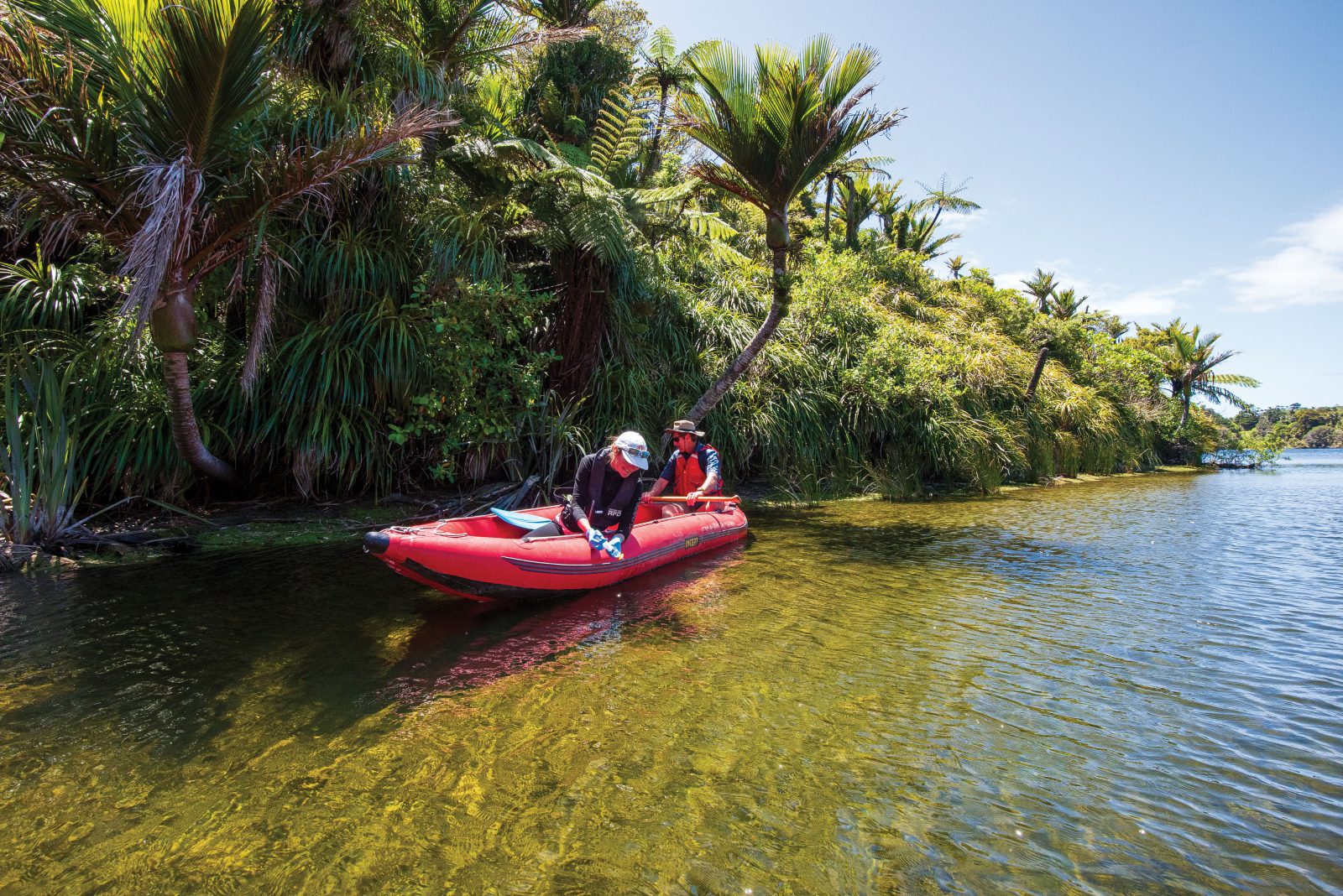
I ponder all this as I watch freshwater scientist Craig Allen draw samples from Pearl Creek as it dawdles towards Waimea Inlet on Tasman Bay. A pair of kōtare/kingfishers are watching him, too, from a power line above, and it occurs to me that they could be pooing all sorts of false flags into his samples. “That’s right,” agrees Allen. “They’ve probably been catching marine crabs on the mudflats downstream.”
He says snapper DNA in particular turns up in freshwater assays all over the place, and that’s down to fishers throwing frames and guts into the nearest stream. “We even had marlin in one sample.”
Allen is here testing for the presence of giant kōkopu, the biggest of the native galaxiid family most people know better as whitebait. They’re scarce in Tasman nowadays, but if they’re holding on anywhere, Allen thinks it could be here in Pearl Creek. They like sluggish water, and more than two decades of restoration and replanting work has provided the sort of overhanging vegetation the fish prefer to launch their ambush attacks from.
Pearl Creek is also just the sort of habitat, says Trevor James, that eDNA might have been invented for. He leads the freshwater enhancement programmes at Tasman District Council. “There are some environments that we just can’t sample very well with traditional freshwater fish monitoring,” he says, “particularly those that are slow flowing, deeper, weedy, with lots of woody debris.” (By “traditional”, he means spotlighting, trapping with fyke nets or minnow traps, or electric fishing, in which a surveyor wades a stream like a heron, dipping a low-voltage probe that temporarily stuns any nearby fish. It’s laborious, fussy work that takes days. By contrast, Allen has his six samples filled in maybe 40 minutes.)
James has worked in freshwater ecology for 30 years, which means he’s watched giant kōkopu vanish in lockstep with the habitat it needs: “We have two major ecological districts in Tasman—the Moutere has lost 95 per cent of its wetlands and the Motueka has lost 98 per cent.”
The council has surveyed the species since 2006 and now, he says, “we’re really concerned. We’re finding them at fewer and fewer sites with traditional methods. I think the population’s got to a size where it’s just not able to support itself.” He worries, too, about the brown mudfish, a nondescript, scaleless sliver of a thing named for the fact that it can survive in damp soil for months. They were once common in irrigation ditches across Tasman’s Waimea floodplain, but are now known only from Mangarākau, near the tip of the South Island West Coast. “But we’ve only just started in recent years with eDNA,” offers James, “so we might find a few more yet.”

In the end, just one of the six samples from Pearl Creek held a faint whiff of giant kōkopu DNA, hinting, says James, that “there may be the odd individual hanging in there”.
Revelations elsewhere keep him optimistic: in March 2021, Fiordland College students took samples from the Ewe Burn, just north of Te Anau. Analysis confirmed they’d found a new population of the Gollum galaxias, a fish named after that other bug-eyed, skulking water-dweller. Later that year, eDNA came through again: the Clutha flathead is a critically endangered galaxiid, eaten almost to extinction by introduced trout. By 2015, their entire known global range occupied 12 hectares, but a Department of Conservation eDNA survey found another population—possibly in the hundreds—in Tyre Gully, in the upper Cardrona Valley, where a waterfall had kept trout at bay.
For all its triumphs, James stresses there are important caveats around eDNA: it can tell you only about the presence or absence of something—and, as with the snapper and kangaroo, the presence of DNA doesn’t necessarily mean its owner was alive at the time. It can’t tell you how many of that something there are, or whether they’re male or female, nor how old they are—the sort of stuff an ecologist really needs to know about a population. Some organisms shed DNA more readily than others; some DNA is more resilient, therefore more conspicuous. That can slip a sneaky bias into survey results. It’s important to remember, too, says James, that the DNA you collect at a stream mouth might have travelled tens, maybe even hundreds of kilometres on the current to that point. For that reason, he says, it will only augment traditional hands-on survey methods.
[Chapter Break]
Other times, eDNA is useful for telling you what’s not there. In 2018, Neil Gemmell, a geneticist at the University of Otago, took 250 samples from Loch Ness—some as deep as 200 metres—then combed them for essence of monster. “We didn’t find it,” he told media. “Really, it was just a great way to showcase the power of eDNA. And, you know, we might’ve found a monster.”
Those long, chilly days on the water yielded a list of 3000 species, give or take: “Tiny little things like nematodes, roundworms, rotifers, phytoplankton, small crustaceans, those sorts of things. And then, of course, all the fish we expected to find—salmon, pike, eels, sticklebacks.” He also found dogs, sheep, cattle and badgers, but no plesiosaurs. Nor was there any sign of two other monster suspects—sturgeon or catfish.
Here in Aotearoa, a dedicated band is still searching for the South Island kōkako—nominally extinct till proven otherwise. The last accepted sighting was in 1967, and the bird was declared lost in 2008. But a reliable account from near Murchison in 2007 raised sufficient doubt to resurrect the kōkako: it’s now listed as “data deficient”.
Inger Perkins is the manager of the South Island Kōkako Charitable Trust. She says the group hopes eDNA sampling might prove the bird’s continued existence, but standard water sampling has so far drawn a blank. Proof, she says, may yet be in the wind. “Perhaps installing a small portable air filter in the middle of the forest might be our next step.”

Beyond mythbusting, the applications for biosecurity are legion, and pest managers are already using eDNA along the Waikato River to get a handle on the spread of the Asian gold clam, a calamitous invasive bivalve costing authorities a fortune across North America and Europe. In Tasman, eDNA helps the regional council track the spread of water celery and Himalayan balsam—the stuff of biosecurity nightmares.
“It’s pretty cool, right?” says Jaret Bilewitch. “It’s kind of like CSI on television. It borders on forensics, and this whole idea of detecting something without being able to see it is quite attractive.” Just the same, as a molecular biologist he knows there are hard limits to what it can do, especially at sea.
At the National Institute of Water and Atmospheric Research (NIWA), Bilewitch leads the development of eDNA tools for finding biodiversity—wanted and unwanted—and points out that the sea is a very different sampling environment from a stream or a sewage pond. Typically, he says, biosecurity teams will use the community profiling approach first, to get a picture of the broader community of organisms at a site. “Then we analyse the list of species that comes out of that, and ask: ‘Are there any red flags here that we need to be concerned about?’”

A few years ago, broad-scale eDNA sampling in Waitematā Harbour turned up a very big red flag: a single-celled pathogen called Bonamia. For commercial oyster fisheries like those at Bluff and Rakiura, a particular strain of this pathogen, called Bonamia ostreae, is the zombie apocalypse. It was first detected in 2015, in the Marlborough Sounds, where it soon wiped out entire oyster farms. Had the apocalypse spread north? It was a tense wait while NIWA ran their own, more sensitive, tests over the sample.
But it turns out we have our own endemic strain of Bonamia, and the first test couldn’t tell it apart from the nasty one. “We were able to reassure people that it wasn’t the one they needed to worry about,” Bilewitch says.
That scare highlighted the first fundament of biosecurity: in order to know what’s invasive, you first need to know what’s native, and marine organisms have always been the Cinderellas of Aotearoa’s biodiversity. NIWA holds the most comprehensive collection of marine invertebrates in the country, and hundreds of new species are added every year. But there are too few taxonomists to describe them all. The discipline has been hobbled for decades by mean funding and an image problem: young graduates just don’t think taxonomy is sexy. That in turn means there are often no reference DNA barcodes for Bilewitch to turn to. “We need taxonomists to tell native from invasive; we need the baseline information they provide. It just keeps coming up over and over again.”

Case in point: In 2021, ecologist Jack Warden noticed a strange new seaweed in Blind Bay, on Aotea/Great Barrier Island [see Issue 177]. When staff of the Ministry for Primary Industries (MPI) saw his photos, they knew right away the news wasn’t good—the pictures showed that Caulerpa brachypus, already a scourge overseas, had arrived in Aotearoa. It fell to NIWA to formally identify the algae. The job was tougher for the fact that Aotearoa also has its own native Caulerpa species; Bilewitch says eDNA was no help because the caulerpas weren’t on the database. Despite the high stakes—this is a seaweed with the potential to smother ecosystems right along our coastline—“We really don’t know how many species there are, or as much about their distribution as we’d like.”
So eyeballs will always have their place. Divers, remote-operated vehicles and remote cameras, Bilewitch says, are still NIWA’s go-to surveillance and monitoring tools: “We’ve taken a pretty cautionary approach to eDNA, because there have been instances of obvious species getting missed altogether, or detections that turned out to be wrong. People’s expectations and understanding of what it can and can’t do have been a bit unrealistic.”
[Chapter Break]
For Mere Tamanui, eDNA is a way to stitch people and place back together. Surprise a community with a taonga species they thought was long gone, or an invasive one that’s just arrived, and you activate kaitiakitanga, she says, and invite the people to assert their tino rangatiratanga. “It gives us a push forward,” the Tolaga Bay freshwater ecologist says.
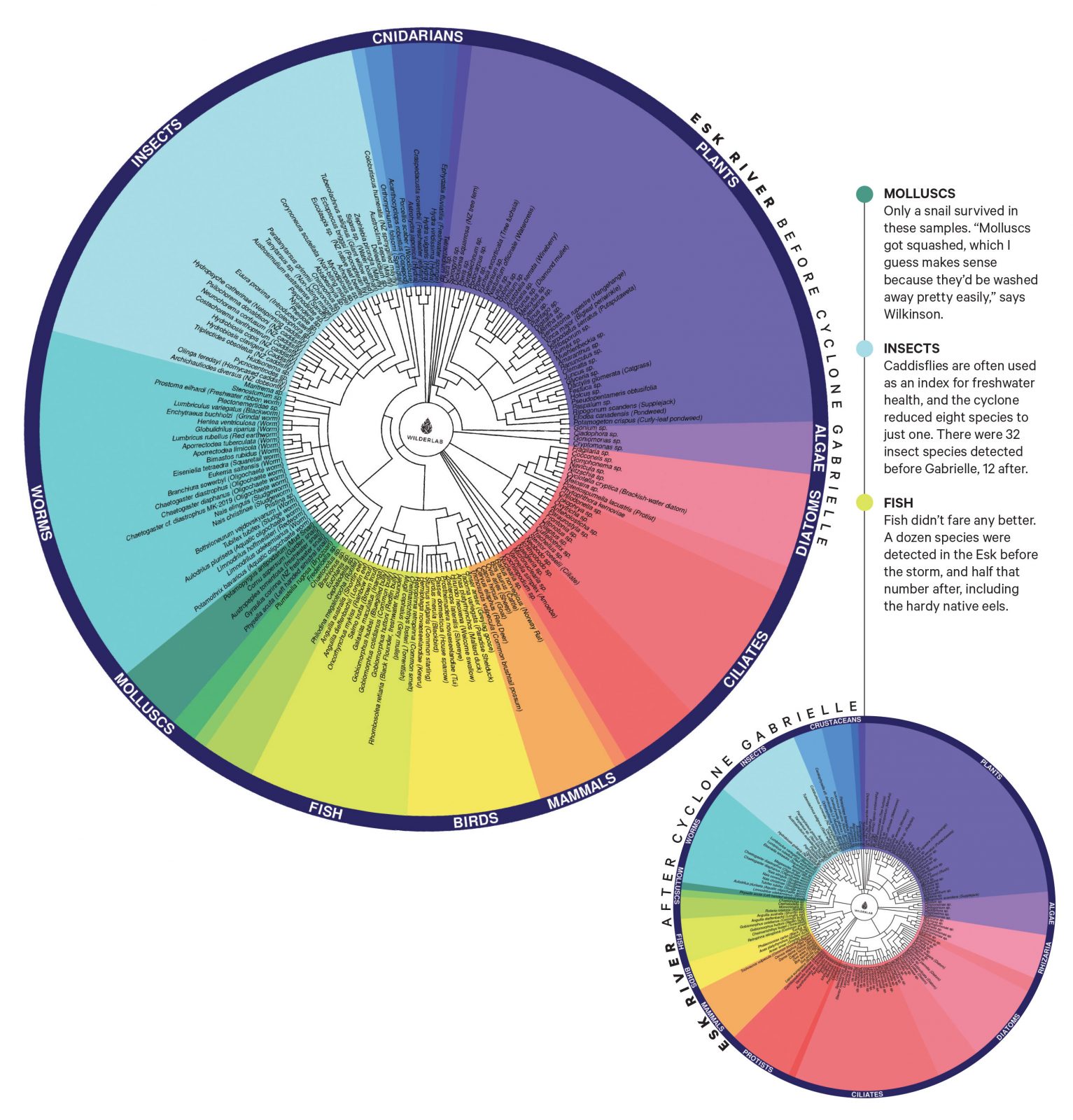
For more than 10 years, Tamanui (Te Aitanga-a-Hauiti, Ngāriki Kaiputahi, Te Aitanga a Mahaki, Ngāti Porou) has been watching her region’s waterways fall apart. By the time Cyclone Gabrielle hit in February last year, she was almost past feeling it. Yet she was out there, hanging a sampling kit off the Ūawa bridge 24 hours ahead of the storm, determined to get a “before” result as a disaster-impact baseline. She took her sample from the spot where the nannies used to fish: knitting Western science with mātauranga.
This summer, a year on, Tamanui and the groups she’s trained are testing a stretch of the same river, looking for kākahi, the precious mussels that filter muck from the water, as well as the cockabullies and mullet that give the mussels’ eggs hitchhike rides and will help the species recover. They have already tested here with no luck. But the floodwaters have receded a bit further now. And miracles happen: recently, eDNA threw up kōaro, or climbing galaxias, in a stretch of water that had been “absolutely ravaged”—by sediment, floodwaters, forestry slash, the diggers that went in to pull the slash out. If she were a fish, Tamanui laughs sadly, she would not be keen on that patch of river.
She tested on a whim, expecting to find nothing much. “That this species is still out there, for us, is absolutely wonderful. It gives us hope.”
Tamanui also appreciates that eDNA is a “safe tool”, in that it can’t spit out the sort of data MPI might use to figure out quota allocations, ie start fishing that species. “If it was giving out population [levels] it would be a different story for us.” To her the technology is fish-friendly, kid-friendly, and it saves masses of time and energy, allowing her already-stretched community to focus on the most important work.
While Tamanui works on the awa, Graeme Atkins (Ngāti Porou, Rongomaiwahine) is using eDNA to inform pest control—1080 drops and deer culling—in the region’s rugged Raukūmara forest. Thirty years ago, it was a “garden of Eden”, the taiao kaimahi/Department of Conservation (DOC) ranger says. “It’s been eaten to death, basically—too many deer and possums.”
Last year, heading back in after a poison drop, the first thing Atkins noticed was the birdsong, and the coprosma fruit all over the ground. Such a feast would previously have been scoffed by possums and rats. But not every predator was wiped out. His team swabbed dozens of fresh deer droppings and sent the samples off for analysis. The idea was to pin down a genetic profile of the specific deer that survived the drops. When the team carry out annual eDNA sampling in these same spots, they hope to be able to discern whether new deer are arriving, or that remnant population is breeding.
Atkins loves eDNA. A born communicator, he sees its enormous potential for getting communities on board with critical but controversial work like the 1080 programme. There might be Archey’s frogs in this forest, he says, or short-tailed bats. Using trail cameras, they’ve already found kiwi where DOC didn’t know there were any. Friends of his explored the forest in the early 70s and found abandoned kākāpō bowls—the tracks and arenas that males scrape out and meticulously maintain. “Those things were there once. So what else? Using eDNA we could find so much more.” The only roadblock, for him, is the expense of testing.
Tamanui’s eDNA work is funded by the Environmental Protection Authority (EPA), which since 2020 has been distributing Wilderlab’s testing kits and knowhow to Māori communities around the country. Crucially, in this scheme the data is held by mana whenua.


This level of data sovereignty and community involvement is not a given. Phillip Wilcox (Ngāti Kahungunu ki Wairoa, Rongomaiwahine, Ngāti Rakaipaaka) is an associate professor and kaikōkiri Māori—a mentor and advocate—for Māori students at Otago University’s genetics teaching programme. As far as he’s concerned, the science community should have talked eDNA through long before anyone dipped the first syringe.
To Māori, he explains, DNA is a biological embodiment of whakapapa—the woven universe of threads that connect people, creatures and places, all the way back to the atua. As such it’s tapu, and it’s protected by tikanga. “That’s not really acknowledged, so the way these technologies are rolled out can be just another negative colonisation experience, coming as they do on top of a long history of them.” Wilcox points to Article 2 of the Treaty of Waitangi, the clause that assures Māori of rangatiratanga—governance—over taonga. Indigenous species, he points out, are taonga. Therefore, he says, their genetic essence—their DNA sequences— are also taonga. Yet he’s found that scientists, if they consult Māori at all, often do so after the liberty has already been taken: “They get the samples, get the sequencing under way, and then ask us, ‘Hey, how do you feel about this?’ This is about more than intellectual property. It’s undermining Māori rights—it’s undermining the way I see the world.”
Wilcox has spent 20 years developing tikanga—informed guidelines and frameworks—for geneticists to follow. These request that researchers consult Māori first, so that interests can be safeguarded and that communities might even be involved in the work. “Those practices have not been widely adopted,” he laments.
In Aotearoa, Wilcox suspects, people don’t ask first because they fear the answer. But, he explains, “tikanga doesn’t mean ‘no’. It’s just a different way of doing things. So don’t fear it—run with it. Our people know what they’re doing, and we’ll collaborate if there’s a net benefit for us.” With the work done under the EPA scheme, though iwi and hapū own the data, many choose to share their results (you can check out an interactive map at wilderlab.co.nz/explore).
[Chapter Break]
At home in Auckland, Shaun Lee knew he hadn’t found his grayling before he even opened the email. “If it had been a positive result,” he says, “Wilderlab would’ve called me.”
The nil result from Lake Alice joins blanks from Lake Marian, in the Darren Mountains of Fiordland, and the Freshwater River, on Rakiura. But his quest isn’t over: “I’m going to start looking at some of the lakes in the Top of the South.”